COVID-19¶
General information¶
Please see the Swedish COVID-19 Data Portal for the latest information regarding Swedish efforts in COVID-19 research, including data generating facilities. Also see the European COVID-19 Data Portal and Horizon 2020 guidelines regarding COVID-19 for useful information on European level.
Data Life Cycle¶
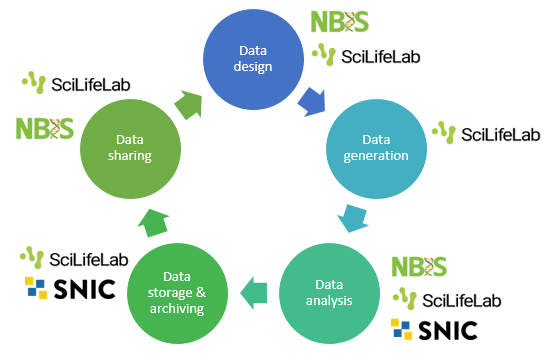
The data life cycle is typically divided into design, generation, analysis, storage & archiving, and sharing. Below you will find information about standards and infrastructure resources available during these phases.
Data design¶
During this phase you plan for wich data is needed to answer your research question. High quality science is often only possible if the resource facilities you intend to use gets involved already in the planning phase of a project. Consultation and advice regarding data management planning, data generation and data analysis are offered by NBIS and SciLifeLab.
It is wise to write a data management plan, using either a tool provided by your university or DS wizard.
Also, some resources have specific application periods and thus needs to be contacted well in advance. If your project includes sensitive human data, note that there are ethical and legal issues that you have to consider, such as apply for an ethics approval and report the data processing to your Data Protection Officer. See the page on Sensitive data for more information.
Data generation¶
SciLifeLab National Genomics Infrastructure (NGI) provide a wide range of sequencing technologies and can offer state-of-the-art solutions for many different types of COVID-19 sequencing projects. Chemical proteomics & proteogenomics and BioMS offers mass spectrometry support. For a complete list please visit Swedish COVID-19 Data Portal.
Data analysis¶
- NBIS (National Bioinformatics Infrastructure Sweden) national research infrastructure offers bioinformatic support in various forms for a wide range of areas including NGS, proteomics, metabolomics and biostatistics.
- SNIC (Swedish National Infrastructure for Computing) national research infrastructure makes available large scale high performance computing resources. Apply for Small, Medium, Large or Sensitive data allocation, depending on size and type of project.
Data storage and archiving¶
After the project is finished, the data needs to be stored in a backed-up fashion at least for 10 years, and for as long as the data is of scientific value. After this time, some of the data should be archived and some can be disposed. It is best to contact your university Research Data Office for information about the procedures for this.
SNIC offers storage for small and medium-sized datasets. In the future also large-sized storage will be offered.
Data sharing¶
The guidelines in all subsections regarding COVID-19 has been adapted from the Research Data Alliance 5th release of the COVID-19 Data Sharing Recommendations & Guidelines.
In general:
- Think early about systematic naming of filenames. Not thinking about it early enough is often the cause of a lot of extra work when the data is not stored in a database and researchers have to rename a large number of files manually at a later stage.
- Document the computing time and resources required for data processing. This could help other researchers to assess the time and resources required for the pipeline, therefore to decide whether it is feasible to proceed with the local resources available.
- When selecting a repository for submission of the data, priority should be given to domain-specific repositories over generic (e.g. institutional) repositories. Domain-specific repositories are easier to find, and often have better visualization and selection facilities for re-users of the data.
- The repositories listed for deposition are also prime locations for locating existing data. Many now have dedicated sections for new as well as pre-existing data relevant to Covid19 research.
The following subsections contain guidelines adressing specific covid-19 data types and resources: